Computational models have been important for studying the connection between form and function involving fluid flow. However, understanding evolution of such structures requires understanding how variation in morphology and kinematics impacts functional performance. Exploring this functional space through simulations is computationally expensive and difficult.
In a collaboration with Dr. Yanyan He (Mathematics, New Mexico Tech), we are using uncertainty quantification to both assess the impact of variation on functional performance and make predictions about the patterns of form that might be observed in existing organismal diversity.
Applying sensitivity analyses to the quantified output variation allows us to understand:
– what input parameters are very sensitive, meaning small changes lead to big changes in output, and likely to be highly conserved.
– what input parameters are not sensitive, meaning changes have no consequences to output performance, and are likely free to diversify.
– what input parameters interact synergistically.
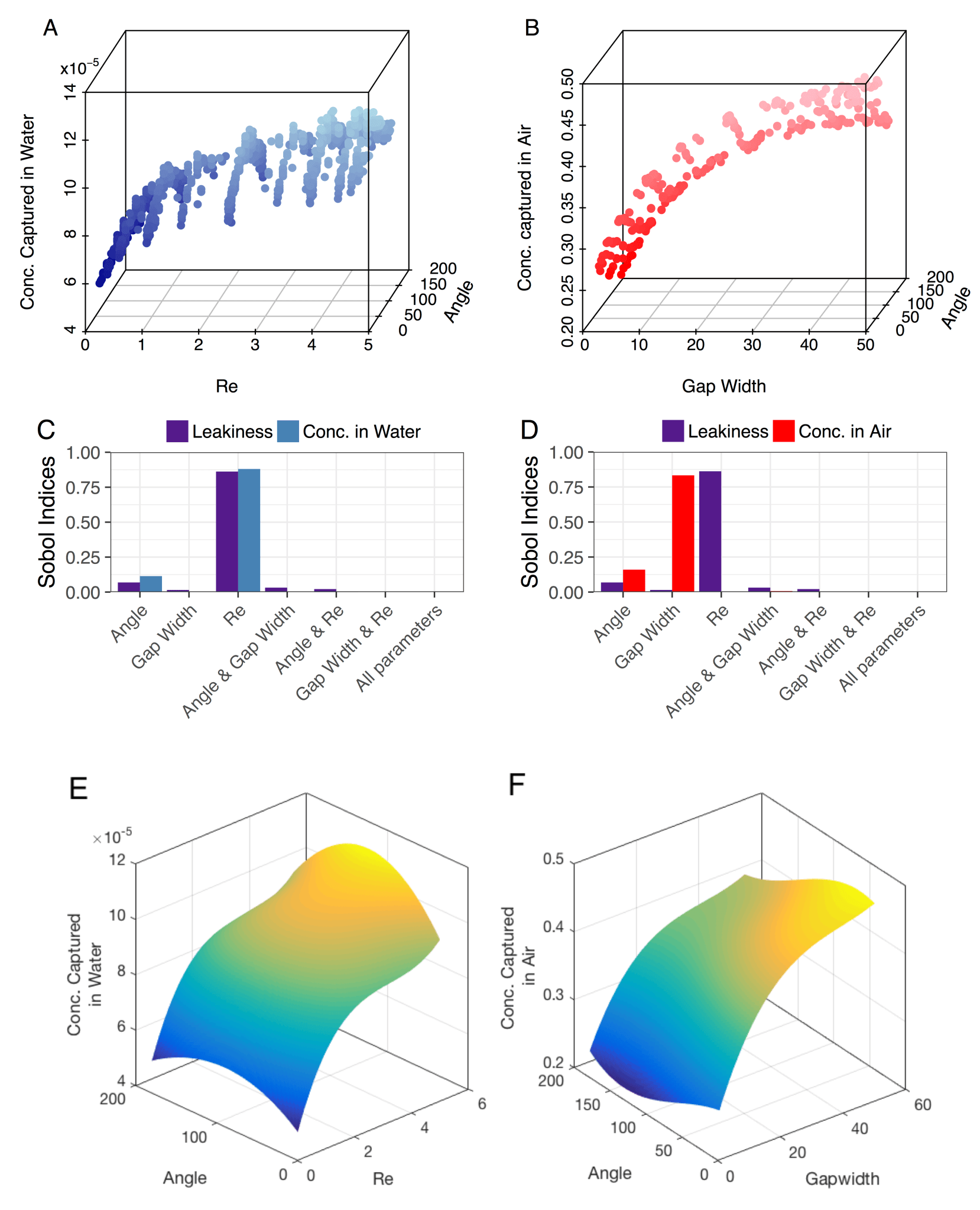
Fig. 5 from Waldrop et al. 2018 showing the concentration of odor captured in air and water against three input parameters for a theoretical chemosensory hair array. The parameter space is sampled using generalized polynomial chaos (A & B) and the sensitivity analyses (C & D) were calculated based on the output of full simulations. Surrogate plots are shown in E & F.
Associated Publications
L.D. Waldrop, Y. He, S. Khatri. 2018. What can computational modeling tell us about the diversity of odor-capture structures in the Pancrustacea? Journal of Chemical Ecology https://link.springer.com/article/10.1007/s10886-018-1017-2. Full text.
L.D. Waldrop, Y. He, N. Battista, T. Neary, L.A. Miller. Uncertainty quantification as a method to infer physical constraints on parameters of pumping by valveless, tubular hearts. In prep.